Team members
Hanyu Wang
Muscle dynamics
We are investigating the mechanobiology of muscle. We are interested in how functional muscles are made during development and how they remain functional through the lifespan of an animal.
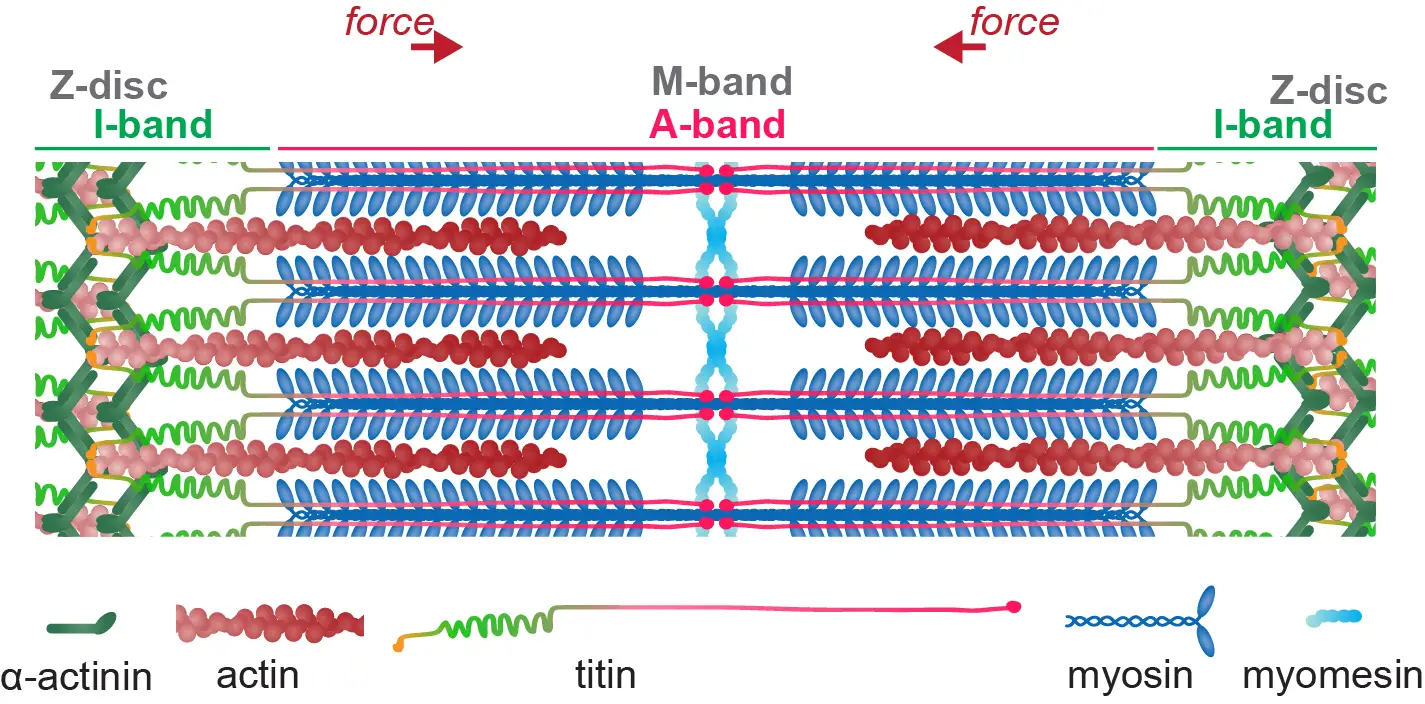
The strength of our group lies in its unique combination of systematic genetics and live in vivo cell biology, with mechano- and structural biology. We are using Drosophila in vivo and human iPSC in vitro models to study the mechanisms how muscle assemble their contractile sarcomeres and how these sarcomeres are serviced in the living animal to remain functional throughout life. Sarcomeres are amongst the largest protein assemblies in animals: they produce high forces and mechanically link to the skeleton to power animal movement. Hence, sarcomeres are a fantastic playground to understand basic principles of biology:
How do thousands of large proteins assemble to build a micro-meter large pseudo-crystalline sarcomere?
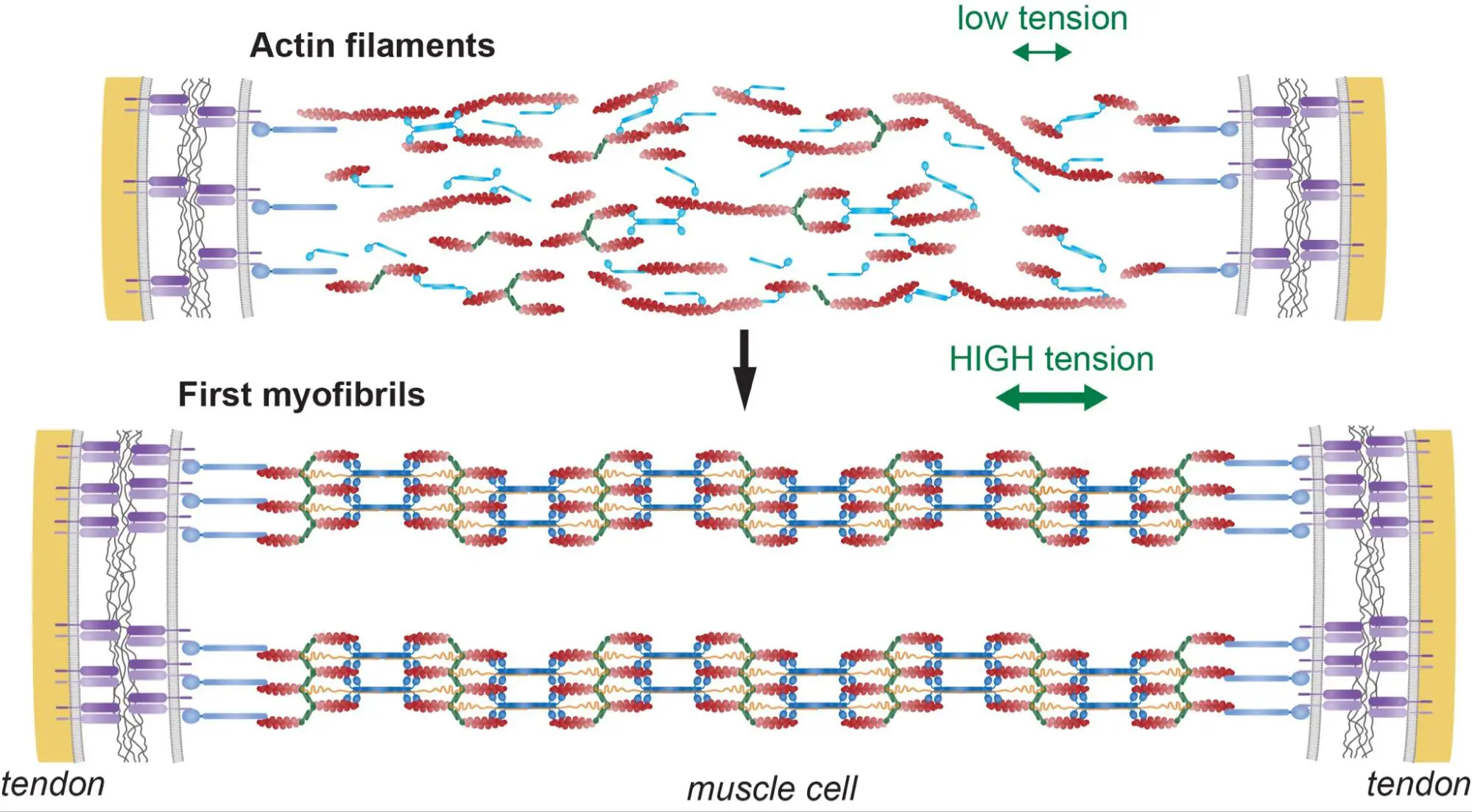
How do thousands of sarcomeres self-assemble to chains that mechanically connect across centi-meter long muscle fibers?
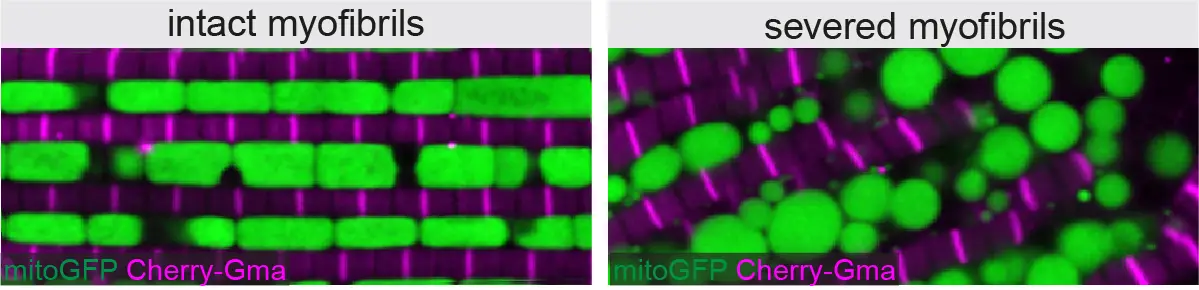
How are sarcomere development and maintenance coordinated with the other physiological requirements of muscle cells including mitochondria biogenesis, T-tubule formation and proteasomal turn-over of damaged proteins throughout life?
Answering these questions will enable us to better understand how muscle are effectively built during development, how they adapt to their different physiological needs (see heart vs. skeletal muscles) and how they remain functional for our entire life.
Publications
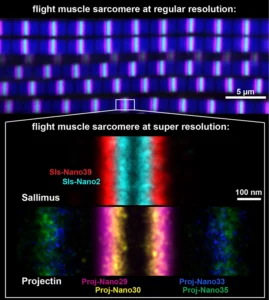
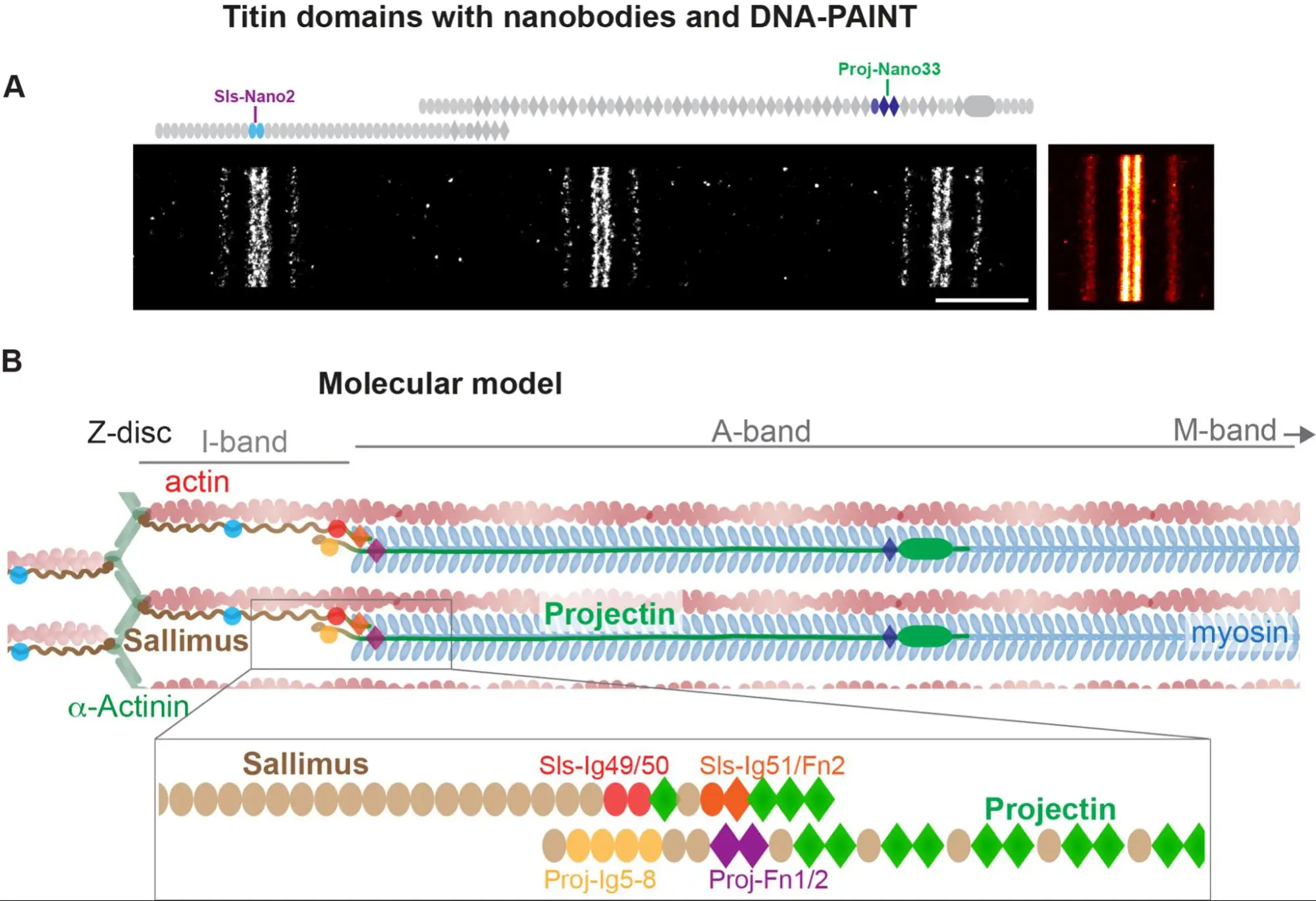
Nanobodies combined with DNA-PAINT super-resolution reveal a staggered titin nanoarchitecture in flight muscles
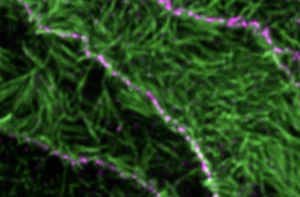
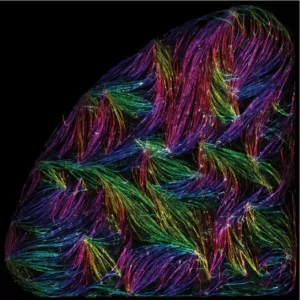
Tension-driven multi-scale self-organisation in human iPSC-derived muscle fibers
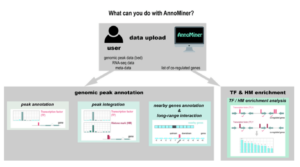
AnnoMiner is a new web-tool to integrate epigenetics, transcription factor occupancy and transcriptomics data to predict transcriptional regulators
Myofibril and mitochondria morphogenesis are coordinated by a mechanical feedback mechanism in muscle
The Hippo pathway controls myofibril assembly and muscle fiber growth by regulating sarcomeric gene expression
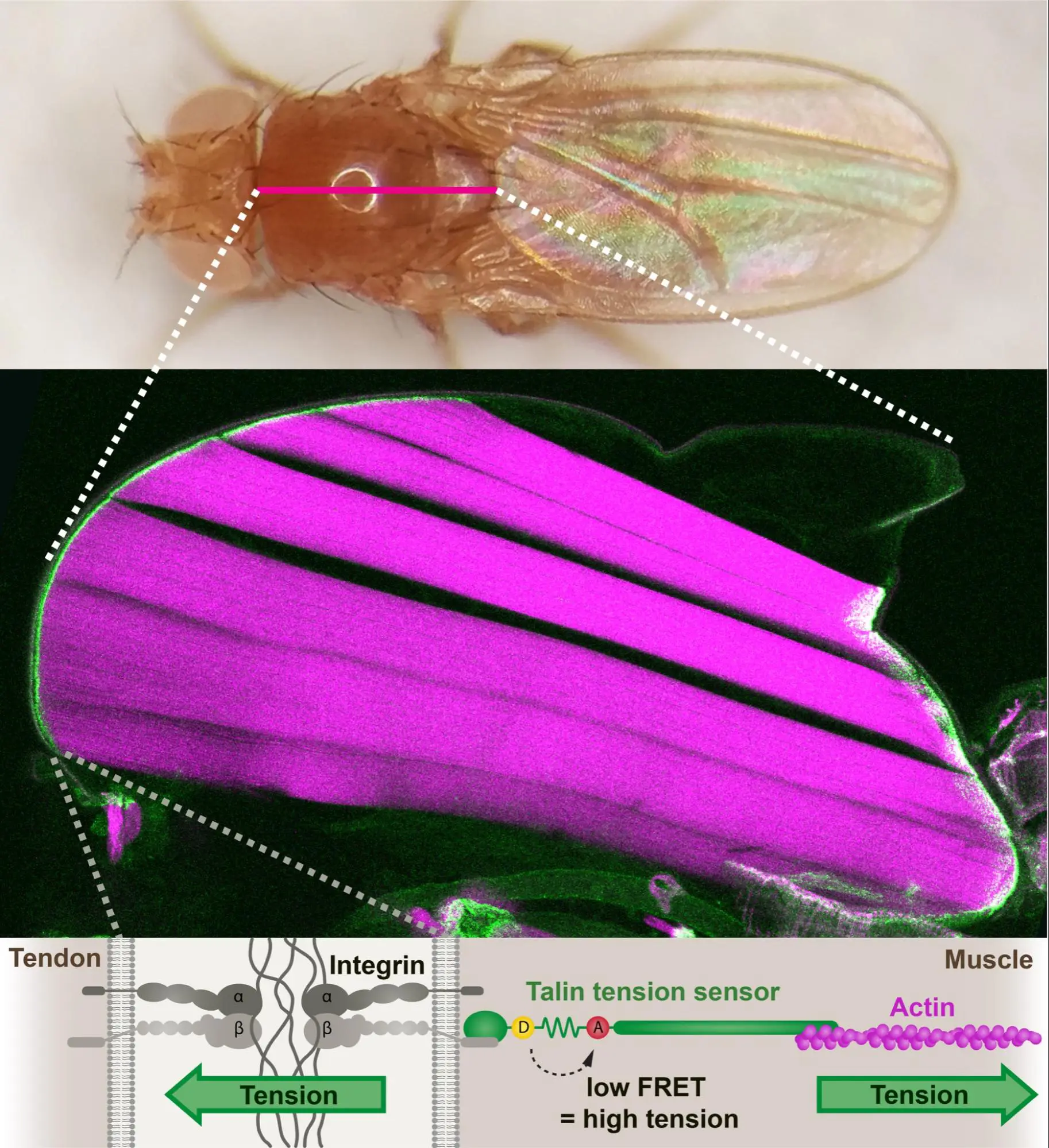
A small proportion of Talin molecules transmit forces at developing muscle attachments in vivo
A nanobody toolbox to investigate localisation and dynamics of Drosophila titins and other key sarcomeric proteins
Nanobodies combined with DNA-PAINT super-resolution reveal a staggered titin nanoarchitecture in flight muscles
Tension-driven multi-scale self-organisation in human iPSC-derived muscle fibers
Tagging Drosophila Proteins with Genetically Encoded Fluorophores
Fly Cell Atlas: A single-nucleus transcriptomic atlas of the adult fruit fly
Mechanobiology of muscle and myofibril morphogenesis
AnnoMiner is a new web-tool to integrate epigenetics, transcription factor occupancy and transcriptomics data to predict transcriptional regulators
Myofibril and mitochondria morphogenesis are coordinated by a mechanical feedback mechanism in muscle
The Hippo pathway controls myofibril assembly and muscle fiber growth by regulating sarcomeric gene expression
A small proportion of Talin molecules transmit forces at developing muscle attachments in vivo
A transcriptomics resource reveals a transcriptional transition during ordered sarcomere morphogenesis in flight muscle.
Ordering of myosin II filaments driven by mechanical forces: experiments and theory.
Polarization-resolved microscopy reveals a muscle myosin motor-independent mechanism of molecular actin ordering during sarcomere maturation.
In Vivo Imaging of Muscle-tendon Morphogenesis in Drosophila Pupae.
Gene Tagging Strategies To Assess Protein Expression, Localization, and Function in Drosophila.
Mechanical tension and spontaneous muscle twitching precede the formation of cross-striated muscle in vivo.
A Guide to Genome-Wide In Vivo RNAi Applications in Drosophila.
A genome-wide resource for the analysis of protein localisation in Drosophila.
Slit cleavage is essential for producing an active, stable, non-diffusible short-range signal that guides muscle migration.
The RNA-binding protein Arrest (Bruno) regulates alternative splicing to enable myofibril maturation in Drosophila flight muscle.
Tension and force-resistant attachment are essential for myofibrillogenesis in Drosophila flight muscle
Ret rescues mitochondrial morphology and muscle degeneration of Drosophila Pink1 mutants
Transcriptional regulation and alternative splicing cooperate in muscle fiber-type specification in flies and mammals
A simple protocol to efficiently engineer the Drosophila genome by TALENs
Spalt mediates an evolutionary conserved switch to fibrillar muscle fate in insects
The Drosophila blood brain barrier is maintained by GPCR-dependent dynamic actin structures.
Systematic genetic analysis of muscle morphogenesis and function in Drosophila
Three-dimensional reconstruction and segmentation of intact Drosophila by ultramicroscopy.
In vivo RNAi rescue in Drosophila melanogaster with genomic transgenes from Drosophila pseudoobscura
High-resolution, high-throughput SNP mapping in Drosophila melanogaster
Ultramicroscopy: 3D reconstruction of large microscopical specimens
Positional cloning by fast-track SNP-mapping in Drosophila melanogaster
A genome-wide transgenic RNAi library for conditional gene inactivation in Drosophila.
The transmembrane protein Kon-tiki couples to Dgrip to mediate myotube targeting in Drosophila
The gammaTuRC components Grip75 and Grip128 have an essential microtubule-anchoring function in the Drosophila germline
RhoGEF2 and the formin Dia control the formation of the furrow canal by directed actin assembly during Drosophila cellularisation
Muscle building; mechanisms of myotube guidance and attachment site selection
Gamma-tubulin37C and gamma-tubulin ring complex protein 75 are essential for bicoid RNA localization during drosophila oogenesis
The molecular motor dynein is involved in targeting swallow and bicoid RNA to the anterior pole of Drosophila oocytes.
Oligomerisation of Tube and Pelle leads to nuclear localisation of dorsal
The cellular localization of the murine serine/arginine-rich protein kinase CLK2 is regulated by serine 141 autophosphorylation
News
Join the IBDM for your internship!
Seeking for your Master internship? The IBDM seems like the right place to do it? Check out our offers.
Congratulations to Robert Kelly, Frank Schnorrer, Cédric Maurange, Bianca Habermann and Delphine Delacour!
From AFM-Téléthon Postdoc Fellow to CNRS Researcher: Qiyan Mao is advancing human muscle research through molecular and tissue mechanics approaches.
IBDM Marseille inspires young minds: engaging primary school children on childhood cancer (“Contre le cancer, j’apporte ma pierre”) and interacting with high school students through immersive experiences (DECLICS).
Internal Seminar by Nuno Luis
Join us on 13/07/2023 at 12:30 in Amphi 12 for an exciting talk by Nuno Luis from our Team!
Drosophila sarcomeres visualised at super-resolution with the help of nanobodies and single molecule blinks.
Organizing the organizers
Self-organisation of human muscles in a dish
Human muscle cells self-organise into defined fiber bundles in vitro even without the presence of external cues !
Frank Schnorrer elected as EMBO member
EMBO elects 67 new members and associate members. They join the community of more than 1,900 leading life scientists in Europe and beyond.
We introduce a novel, user-friendly web-based tool ‘AnnoMiner’ to annotate and integrate epigenetic and transcription factor binding data.
By combining genetics in the fruit fly Drosophila with state-of-the-art imaging and deep-learning, researchers at IBDM have found that mitochondria coordinate their formation with myofibril development to match the correct muscle type.
The Hippo pathway controls muscle growth
Schnorrer team and colleagues discovered that a signalling pathway, called the Hippo pathway, is controlling muscle growth during development of Drosophila flight muscles.
European Synergy Grant for Muscle Research
The European Research Council (ERC) awarded one of the rare ERC Synergy Grants to an international consortium of scientists, Frank Schnorrer, Stefan Raunser, Dirk Görlich and Mathias Gautel.
Investigating the mechanobiology of self-organisation in human iPSC-derived skeletal muscles
The successful completion of the project will allow you to answer how mechanical tension is generated inside human myofibers by integrating intracellular molecular forces and extracellular mechanical environment.
Investigating the mechanobiology of self-organisation in human iPSC-derived skeletal muscles
Project title: Investigating the mechanobiology of self-organisation in human iPSC-derived skeletal muscles Type of rotation: M1 (2 months) or M2 (6 months) Supervisors: Qiyan MAO